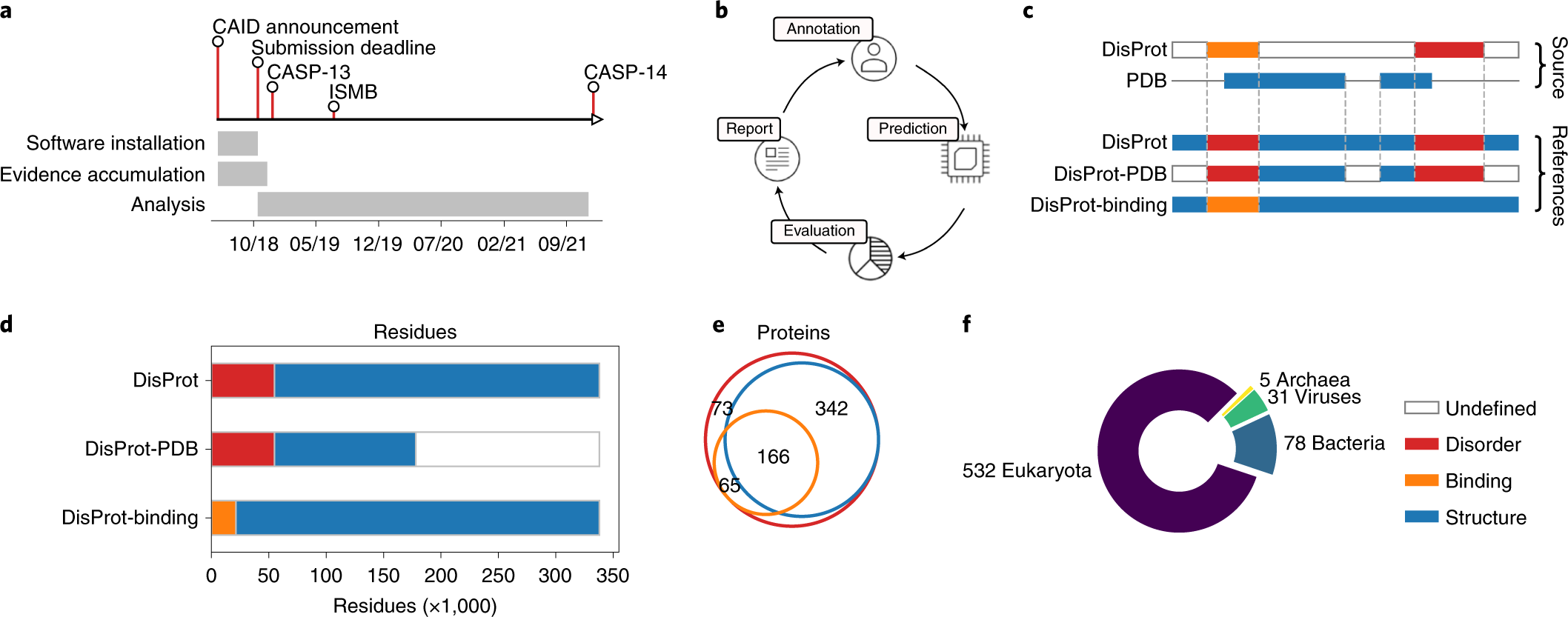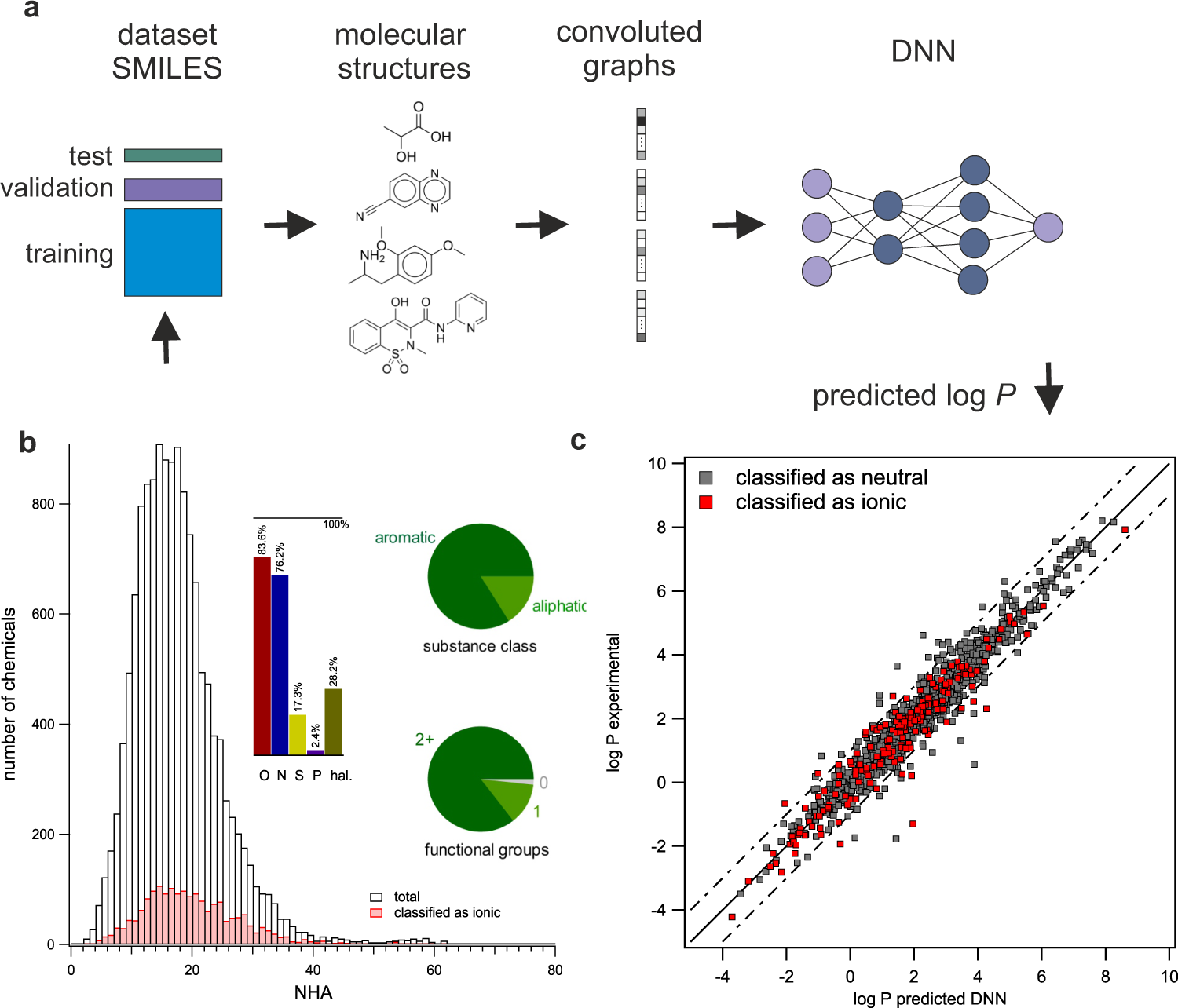
Exploring the octanol–water partition coefficient dataset using deep learning techniques and data augmentation | Communications Chemistry

Collaborative Approach between Explainable Artificial Intelligence and Simplified Chemical Interactions to Explore Active Ligands for Cyclin-Dependent Kinase 2 | ACS Omega

Predicting Power Conversion Efficiency of Organic Photovoltaics: Models and Data Analysis | ACS Omega
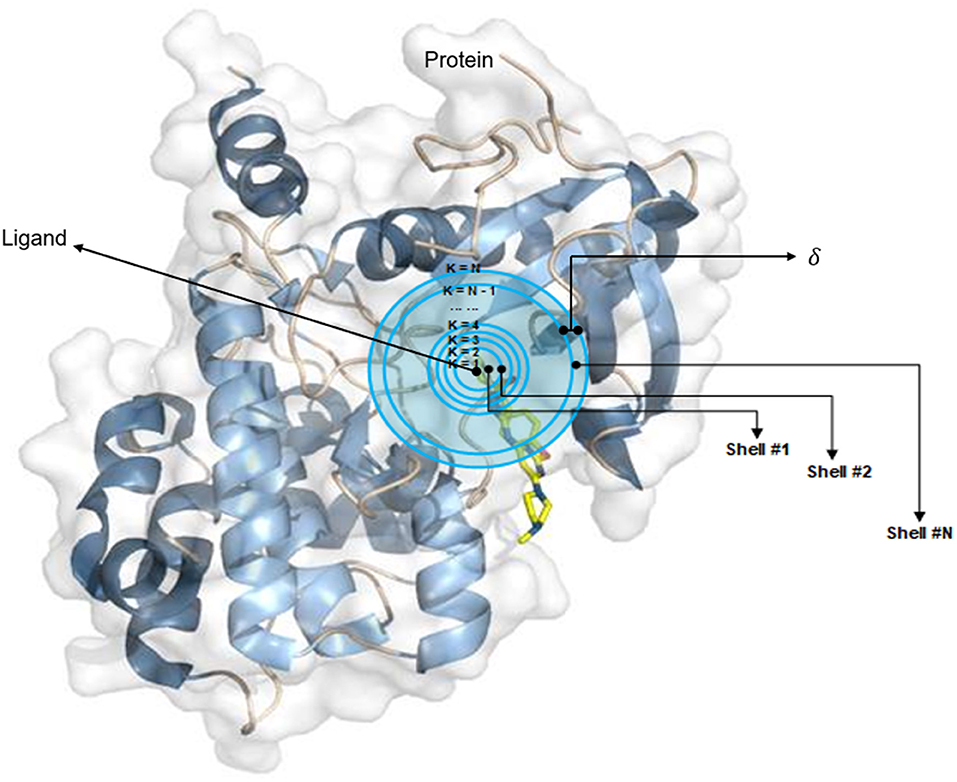
Frontiers | SE-OnionNet: A Convolution Neural Network for Protein–Ligand Binding Affinity Prediction | Genetics

Real-Time Prediction of Equivalent Circulation Density for Horizontal Wells Using Intelligent Machines | ACS Omega
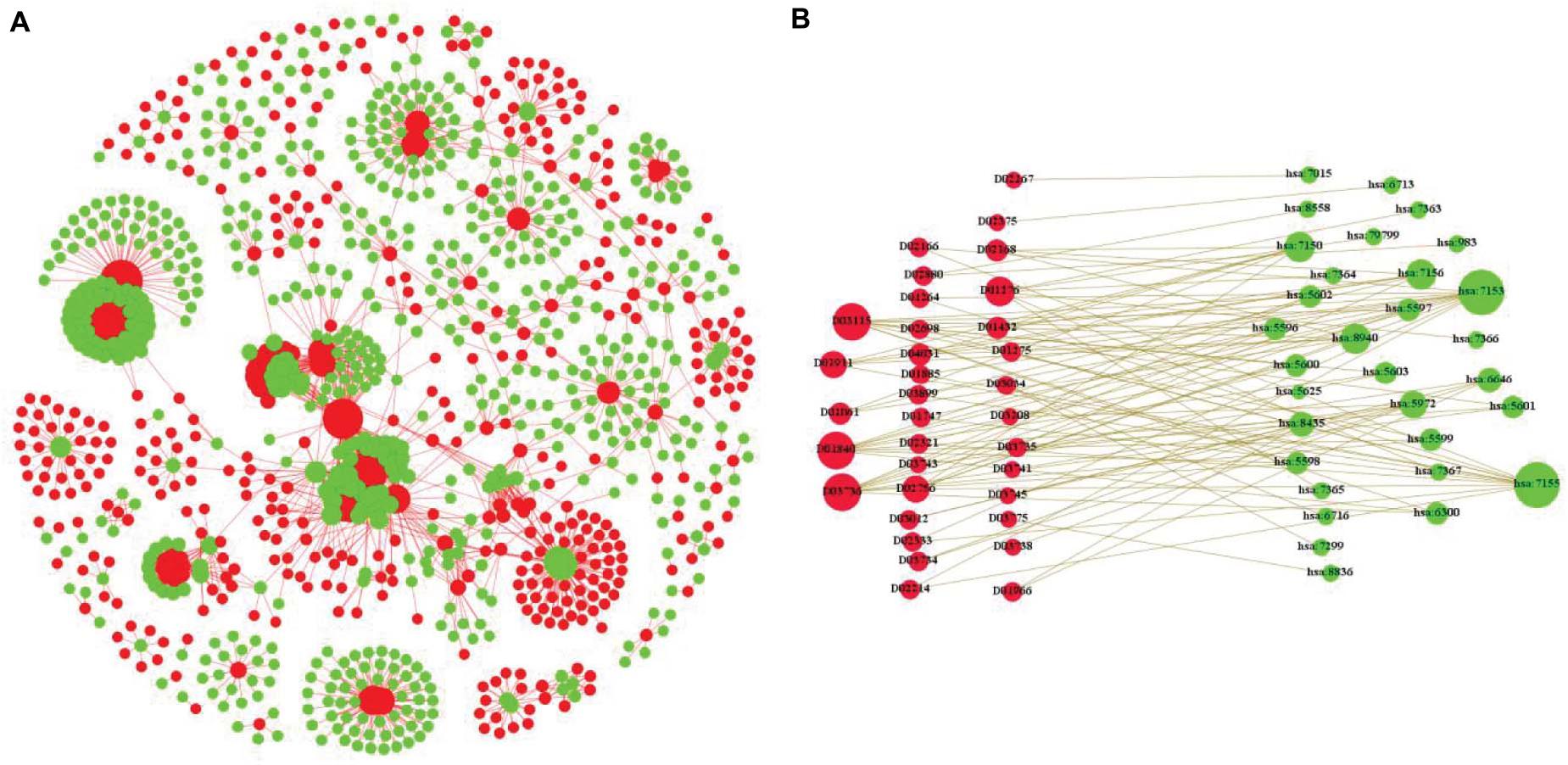
Frontiers | Cascade Deep Forest With Heterogeneous Similarity Measures for Drug–Target Interaction Prediction | Genetics
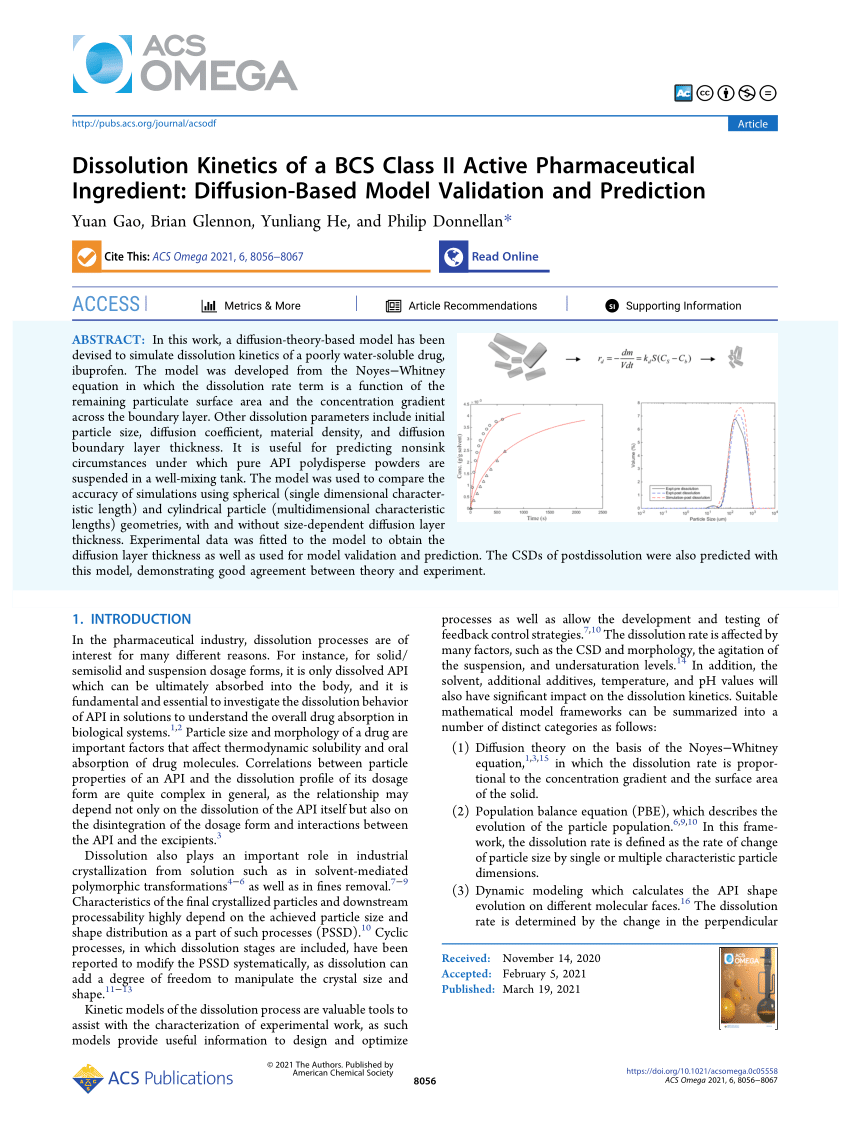
PDF) Dissolution Kinetics of a BCS Class II Active Pharmaceutical Ingredient: Diffusion-Based Model Validation and Prediction